Introduction
Nucleic acid biosensors — devices that use DNA or RNA sequences as recognition elements — are among the most versatile platforms in analytical chemistry. Their applications span from clinical diagnostics and therapeutic drug monitoring to environmental surveillance and fundamental molecular biology research. Two sub-classes are particularly relevant to SPR: DNA hybridization biosensors, which detect complementary target sequences, and aptamer biosensors, which use short, folded nucleic acid sequences to bind small molecules, proteins, or other non-nucleic acid targets with antibody-like affinity.
Both classes share a common challenge: the development pipeline is slow and expensive. DNA hybridization biosensors often require redox-reporter-modified probes for electrochemical detection — a chemically demanding synthesis that must be repeated for every probe variant under evaluation. Aptamer biosensors face similar complexity: the 3D conformation of the aptamer, its surface density, the matrix it operates in, and the geometry of the sensing layer all determine performance, and each variable requires independent optimization. As DeRosa et al. outline in their widely cited aptamer selection review (Nature Reviews Methods Primers, 2023) — from the LADDER Lab at Carleton University — even a successful SELEX campaign leaves open the structural questions that ultimately govern biosensor performance.
SPR offers a way to compress this development timeline. Because SPR is label-free — detecting binding optically through refractive index changes at the sensor surface — it can interrogate nucleic acid interfaces without any chemical modification of the recognition element. This makes SPR uniquely suited as both a screening tool early in development and as an analytical platform in its own right for continuous monitoring applications.
This technical note covers three case studies that together illustrate SPR's role across the full biosensor development arc — from rational probe design to clinical-grade continuous monitoring — followed by a translational section linking bench research to wearable clinical devices:
- Study 1 (Fontaine et al., Analyst 2025): Combined eSPR and electrochemical detection of DNA hybridization show equivalent analytical performance — validating SPR as a rational design tool before redox-reporter synthesis.
- Study 2 (Park et al., JACS 2025): A split-aptamer cross-linked hydrogel film (aptagel) on a plasmonic surface enables real-time, reversible detection of vancomycin in biological fluids — demonstrating the full potential of aptamer-SPR biosensors for continuous clinical monitoring.
- Study 3 (Basini et al., Nano Today 2026): MD simulation guides the rational design of PNA aptamers for cardiac troponin I — achieving picomolar affinity with nuclease resistance, validated by SPR kinetics, replacing months of SELEX with a targeted computational-experimental pipeline.
Study 1 — DNA Hybridization: SPR as a Rational Design Tool
Fontaine, Dauphin, Gaida, Simard, Dauphin-Ducharme · Université de Sherbrooke · Analyst, 2025, 150, 4389–4394 · DOI: 10.1039/D5AN00741K
Experimental Approach
Fontaine et al. employed an eSPR platform that integrates optical SPR and electrochemical detection on the same gold sensor surface. This simultaneous dual-transduction architecture is critical: because both measurements are collected from the same molecular interface at the same time, any performance differences can be attributed to the detection method alone — not to surface variability or sample preparation differences.
Redox-reporter-modified single-stranded DNA probes of increasing lengths were immobilized on the gold sensor surface. The use of probes with systematically varying lengths enabled a direct assessment of how signal magnitude scales with probe geometry — a key design parameter for any DNA biosensor. Complementary target DNA was then introduced at varying concentrations, spanning the dynamic range of the assay, and both SPR and electrochemical signals were recorded simultaneously at each concentration.
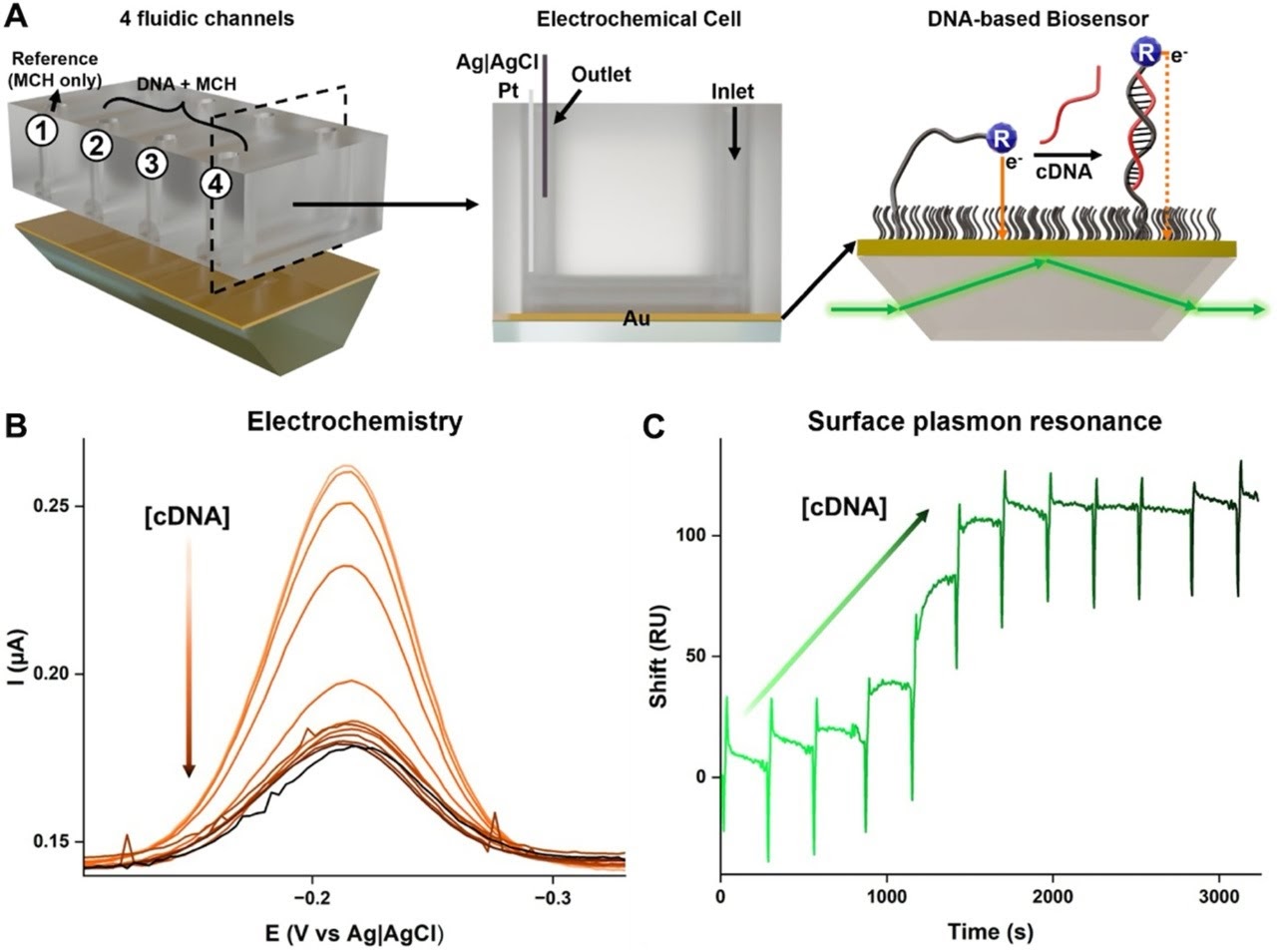
This experimental design is elegant in its simplicity: by combining both measurement modes on a single platform, the researchers eliminated confounding variables and obtained a direct, unambiguous comparison of SPR and electrochemical performance across a systematic probe-length series and a full concentration-response range.
Sigmoidal Dose-Response Curves
Both SPR and electrochemical detection produced sigmoidal concentration-response curves when target DNA concentration was varied across several orders of magnitude. A sigmoidal dose-response profile is the hallmark of a well-behaved affinity biosensor: the signal rises steeply through a sensitive detection window and plateaus at saturation. The fact that both transduction methods reproduced this characteristic shape confirms that the assay behaviour is a property of the DNA hybridization interface — not an artifact of either detection method.
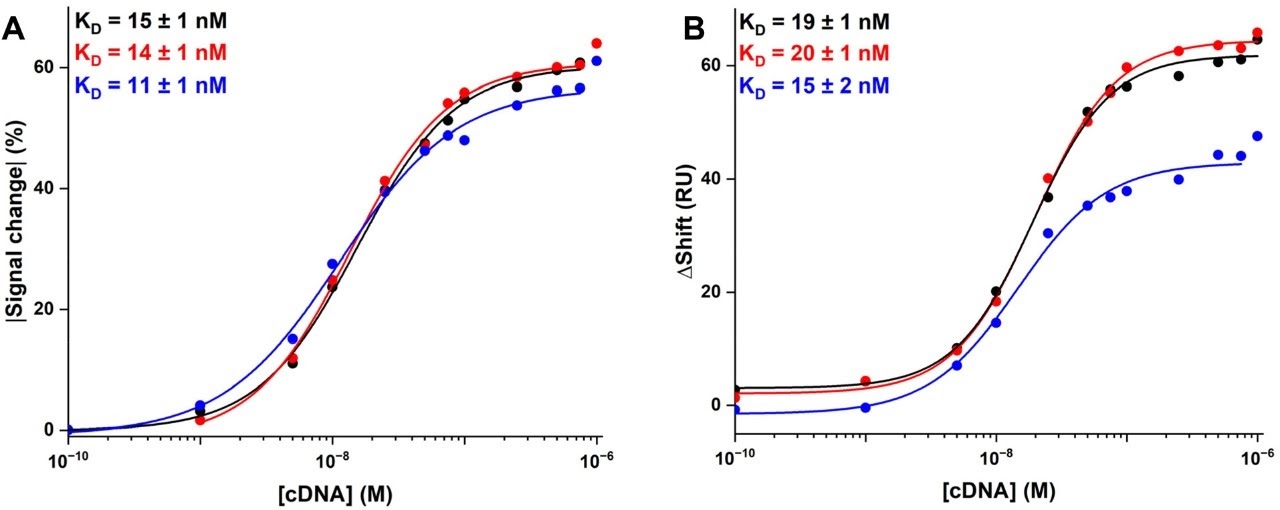
Comparable Limits of Detection
Quantitatively, the limits of detection extracted from the two detection channels were comparable. This is a critical result: it means that SPR is not merely qualitatively similar to electrochemical detection — it achieves equivalent analytical sensitivity. Researchers can therefore use SPR as a quantitative, performance-equivalent screening tool without sacrificing the detection metrics that ultimately matter for the application.
Monotonic Signal Scaling with Probe Length
Signal response varied monotonically as a function of probe length: longer probes produced larger signals in both detection channels. This consistent, predictable relationship between probe geometry and signal magnitude is exactly the kind of design-informing data that makes SPR valuable as an engineering tool. Without the need to synthesize redox-reporter variants for each probe length, researchers can rapidly map the probe-length space using label-free SPR and identify optimal candidates before committing to chemical modification.
Orthogonal Confirmation
The tight agreement between SPR and electrochemical data across the full probe-length series and concentration range provides mutual validation. When two independent transduction methods — one optical, one electrical — interrogate the same molecular interface and produce the same analytical result, the confidence in that result is substantially higher than either measurement alone could provide.
Study 2 — Aptamer Assays: Continuous SPR Monitoring of Vancomycin
Park, Gerber, Santa, Aktug, Hengerer, Clark, Jonas, Dostalek, Sergelen · BioMed X Institute & collaborators · J. Am. Chem. Soc., 2025, 147, 11485–11500 · DOI: 10.1021/jacs.5c01718
Background: Aptamers and the Continuous Monitoring Challenge
Aptamers are short, synthetic nucleic acid sequences — typically 20–80 bases — that fold into defined 3D structures capable of binding a specific target with high affinity and selectivity. Unlike antibodies, aptamers are chemically synthesized (no cell culture required), thermally stable, and can be engineered to undergo conformational changes upon target binding. This structural responsiveness makes them ideal transducers for biosensors: the aptamer itself converts molecular recognition into a measurable signal change.
Vancomycin is a last-resort glycopeptide antibiotic used against multi-drug-resistant Staphylococcus aureus infections. Its therapeutic window is narrow (therapeutic plasma concentrations: ~5–40 mg/L, or roughly 3–27 µM), making real-time drug monitoring during treatment critical to both efficacy and toxicity avoidance. Conventional therapeutic drug monitoring requires blood draws and laboratory analysis — a workflow poorly suited to continuous bedside monitoring. A surface-bound, reversible aptamer biosensor that could directly report vancomycin concentration in biological fluids in real time would be transformative for clinical care.
The Aptagel Platform
Park et al. constructed a new class of SPR-active biosensor layer: a split-aptamer cross-linked hydrogel film, termed an aptagel. The architecture is built from star-shaped 8-arm poly(ethylene glycol) (PEG) macromers cross-linked by split-aptamer pairs — two half-sequences of the vancomycin-binding aptamer, each tethered to the PEG network. In the absence of vancomycin, the network is swollen and relatively open. When vancomycin enters the gel and binds at the aptamer junction, it induces ternary complex formation (split-1 – vancomycin – split-2) that physically contracts the network — increasing its refractive index and decreasing its thickness. Both changes are directly measured by SPR.
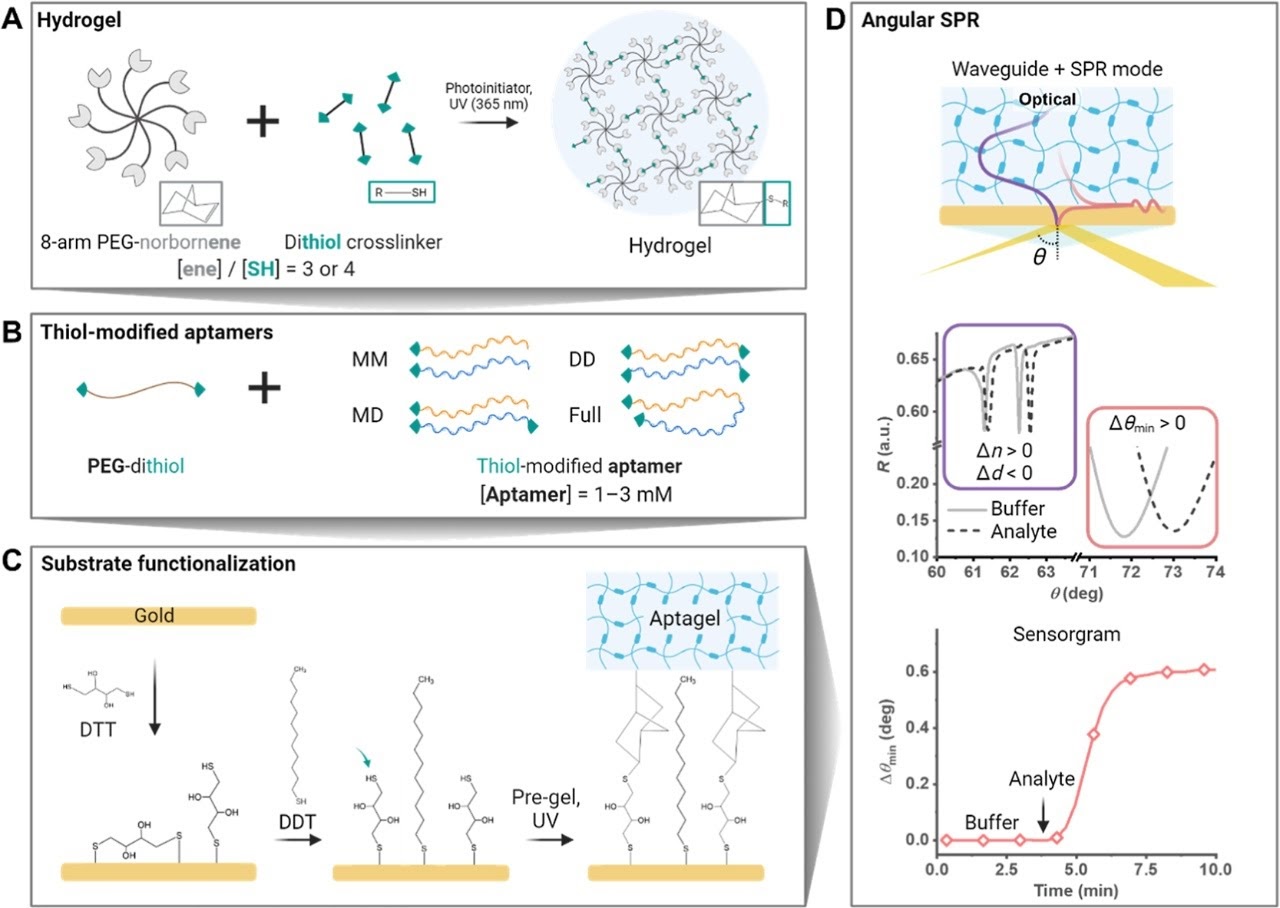
The key design variable was the thiolation strategy used to anchor the split-aptamer halves to the PEG network. Four configurations were evaluated: monothiolated-monothiolated (MM), monothiolated-dithiolated (MD), dithiolated-dithiolated (DD), and a full-sequence aptamer control. The DD configuration — both halves covalently anchored at both ends — proved optimal, delivering the largest and most reversible signal response.
Analytical Performance
| Parameter | Value (SPR mode) | Notes |
|---|---|---|
| Limit of Detection | 160–250 nM | 6–9× improvement over monolayer aptamer |
| LOD (waveguide mode) | 16–30 nM | Enhanced sensitivity via optical waveguide readout |
| Linear dynamic range | Up to 1 mM | Unusually broad for a hydrogel biosensor |
| Full reversibility | < 20 min | Complete signal return without regeneration buffer |
| Stability in diluted plasma | 5 weeks at 37°C | CV = 2.4% |
| Stability in undiluted horse serum | 24 h (90% signal retention) | CV = 2.5% |
| Response time | ~3 min | Both association and dissociation |
| Selectivity | High | <3σ for bacitracin, rifampicin, ampicillin, glucose |
Table 1. Key analytical performance parameters for the vancomycin aptagel biosensor (Park et al., JACS 2025).
The clinical relevance of these numbers is significant. The therapeutic plasma window for vancomycin is approximately 3–27 µM (5–40 mg/L). An SPR LOD of 160–250 nM places the sensor well below the therapeutic range, meaning the assay has ample sensitivity headroom for real patient samples even after dilution. The 5-week stability in diluted plasma at body temperature directly addresses one of the key barriers to implantable or indwelling biosensor deployment.
Why the 3D Hydrogel Outperforms a 2D Monolayer
The dramatic sensitivity improvement over monolayer SPR is not simply a matter of more aptamers per unit area. QCM-D analysis revealed that the DD aptagel undergoes a fundamentally different mechanical response upon vancomycin binding: the contraction is rigid and elastic (a large dissipation-to-frequency ratio change), whereas the full-sequence aptamer monolayer shows a viscous, less organised response. This means the 3D network translates molecular binding events into a coherent, macroscopic mechanical signal — amplifying the optical response far beyond what a flat monolayer surface can achieve.
Equally important, the 3D hydrogel environment dramatically improves stability in biological matrices. A conventional aptamer monolayer on gold loses activity within hours in serum due to non-specific adsorption and competitive displacement. The PEG hydrogel matrix is inherently anti-fouling — it resists protein adsorption — and the aptamer sequences embedded within the network are protected from degradation. The result is a sensor that remains functional for weeks rather than hours in clinically relevant media.
Study 3 — PNA Aptamers: Rational Design, Biostability, and Cardiac Biomarker Detection
Basini, Hambli, Sahu, Corradini · Nano Today, 2026, 67, 102957 · DOI: 10.1016/j.nantod.2025.102957
The Biostability Problem
The studies examined so far — DNA hybridization biosensors and aptamer hydrogels — share an underlying vulnerability: they are built on natural nucleic acids. DNA aptamers, however well-optimized in the lab, face rapid degradation in real biological environments. Blood, plasma, and interstitial fluid contain nucleases — enzymes whose sole function is to cleave nucleic acid sequences. A DNA aptamer biosensor that performs brilliantly in buffer may lose all function within minutes of contact with a blood sample. This biostability gap is one of the central barriers separating laboratory biosensor performance from clinical deployment.
Peptide Nucleic Acids (PNAs) address this problem at the chemical level. PNA is a synthetic nucleic acid analogue in which the sugar-phosphate backbone is replaced with a neutral polyamide chain. PNA retains the ability to form Watson-Crick base pairs with complementary DNA, RNA, and other PNA sequences — but the polyamide backbone is completely invisible to nucleases, which require the sugar-phosphate structure to function. The result is a recognition element with aptamer-like selectivity and affinity, but with the biostability required for point-of-care and in vivo deployment.
Target: Cardiac Troponin I
Cardiac troponin I (cTnI) is the gold-standard biomarker for acute myocardial infarction. Elevated cTnI in blood indicates cardiomyocyte damage with high specificity, and its quantification determines both diagnosis and treatment decisions in chest pain patients. The clinical threshold for detection is in the picomolar-to-nanomolar range — demanding extremely high sensitivity from any biosensor platform. SPR, with its ability to detect refractive index changes from sub-nanomolar binding events, is well positioned for this application.
MD Simulation + SPR: A Rational Aptamer Design Pipeline
Basini et al. combined molecular dynamics (MD) simulations with SPR characterization to rationally design and validate PNA aptamer sequences for cTnI detection. This dual approach represents a significant departure from traditional aptamer discovery workflows. Conventional SELEX (Systematic Evolution of Ligands by EXponential enrichment) requires 8–15 iterative rounds of selection and amplification — a process that typically takes weeks to months and produces aptamers whose structural basis for binding is often poorly understood.
By using MD simulation to model aptamer-target interactions computationally before synthesis, the researchers could predict which structural features of the aptamer governed cTnI binding, then design truncated PNA sequences that preserved only the essential elements. SPR then provided the experimental validation: binding kinetics, affinity constants, and a direct comparison between covalent and non-covalent immobilization strategies — all label-free, in real time, on a single sensor surface.
Key Findings
The MD simulations revealed a critical structural insight: it is the nucleobase sequence within the 3′ end of the aptamer — not the hairpin structure as previously assumed — that governs cTnI interaction. This finding alone changes how future aptamer engineering for this target should be approached. It means that structural truncations preserving the 3′ end can maintain picomolar affinity even in significantly shorter sequences — reducing synthesis cost and complexity without sacrificing performance.
| Parameter | PNA Aptamer | DNA Aptamer (reference) |
|---|---|---|
| Binding affinity (Kd) | Picomolar | Picomolar (longer sequence) |
| Sequence length | Shorter than DNA parent | Full-length required |
| DNase I stability | High — backbone resistant | Low — rapidly degraded |
| Key structural element | 3′ end nucleobase sequence | Hairpin (assumed, not confirmed) |
| Immobilization | Covalent & non-covalent validated | Covalent standard |
| POC suitability | High | Limited by biostability |
Table 2. Comparison of PNA vs. DNA aptamer performance for cTnI detection (Basini et al., Nano Today 2026).
Translational Horizon: From Bench to Wearable
Garg, Chupil, L'Heureux-Hache, Lal, Poudineh, Soleymani · McMaster University · University of Waterloo · Stanford University · Nature Reviews Bioengineering, 2025 · DOI: 10.1038/s44222-025-00394-3
If the three case studies above establish what SPR-based nucleic acid biosensors can do in a laboratory setting, this translational section asks the next question: what happens when these principles reach the body? Researchers at McMaster University, University of Waterloo, and Stanford University — including Mahla Poudineh (Aptec Health, University of Waterloo) — have developed the Aptalyzer: a wearable sensor that integrates hydrogel microneedles with electrochemical aptamer-based sensing for continuous, minimally invasive monitoring of insulin in interstitial fluid (ISF).
The Clinical Problem
Approximately 830 million people live with diabetes globally; 200 million require insulin therapy. Despite this, no clinically deployed device can continuously monitor insulin in real time. Current continuous glucose monitors (CGMs) dominate the wearable biosensor market — valued at US$12.6 billion in 2024 — but are fundamentally limited to glucose detection because they rely on enzyme-based electrochemical sensing. Insulin lacks a suitable enzyme, making CGM-style approaches impossible.
The consequences are significant: insulin pump infusion failures go undetected in up to 41.4% of cases because there is no closed-loop sensor to catch them. A continuous insulin monitor would transform diabetes management — closing the loop between dosing and response, reducing hospitalization, and enabling truly personalized therapy.
The Aptalyzer: Aptamers Meet Microneedles
The Aptalyzer solves the enzyme problem by replacing enzymatic detection with electrochemical aptamer-based (EAB) sensing. Redox-labelled aptamers immobilized on electrode surfaces undergo conformational changes upon binding insulin — translating molecular recognition into an electrical current measurable by square wave voltammetry. Critically, this mechanism is not limited to insulin: it is inherently modular, applicable to any analyte for which a structure-switching aptamer can be selected.
To access ISF without conventional blood draws, the Aptalyzer uses hydrogel microneedles — arrays of sub-millimetre needles that painlessly penetrate the outermost skin layer. The hydrogel swells upon insertion, drawing ISF and its dissolved analytes into contact with the aptamer-functionalized electrode surface. The combination solves two problems simultaneously: it provides continuous, real-time fluid access without repeated sampling, and it positions the sensing interface exactly where the analyte is present.
In Vivo Performance
In preclinical rodent studies, the Aptalyzer demonstrated multi-day continuous monitoring capability — successfully tracking dynamic changes in insulin, glucose, and lactate simultaneously in response to injections. Measured concentrations correlated strongly with blood-based assays. The device operated for up to three days before signal degradation occurred, meeting the threshold considered necessary for meaningful clinical utility and approaching the 10-day benchmark set by commercial CGMs.
From Academic Prototype to Commercial Product
The Aptalyzer is not an isolated proof of concept — it represents the leading edge of a broader commercial wave. Nutromics, an Australian medtech company, is building the Lab-on-a-Patch® on the same architectural logic: electrochemical aptamer-based sensing integrated with microneedle sampling in a wearable patch format. Where the Aptalyzer is a research platform optimized for a specific analyte (insulin) in a preclinical setting, the Lab-on-a-Patch® is designed as a generalized continuous diagnostic platform — intended to monitor multiple biomarkers simultaneously in a clinical or consumer health context.
Aptec Health, co-founded by Mahla Poudineh at the University of Waterloo, is pursuing the same trajectory from a different starting point — translating the Aptalyzer platform itself toward clinical deployment for continuous insulin monitoring in people with diabetes.
The convergence is not coincidental. Both platforms derive from the same core insight: EAB sensing is inherently modular, and microneedle sampling solves the access problem without the pain and inconvenience of repeated blood draws. Once the aptamer selection, surface chemistry, and hydrogel matrix are optimized — the work that benchtop SPR is built to accelerate — the sensing architecture transfers directly from bench to patch.
P4SPR Advantages for Nucleic Acid Biosensor Development
No Redox Labels Required
SPR detects hybridization optically. No redox-reporter synthesis is needed for initial assay development — start screening immediately with unmodified probes.
Accelerate Design Cycles
Screen probe lengths, surface densities, and buffer conditions in hours before committing to costly chemical modifications. Fail fast and cheaply — succeed with confidence.
Real-Time Hybridization Kinetics
Watch association and dissociation in real time. Identify optimal probe designs — and eliminate poor performers — before electrochemical validation is ever attempted.
Equivalent Sensitivity
Comparable limits of detection to electrochemical methods mean you are not sacrificing analytical performance for the convenience and speed of label-free SPR screening.
Orthogonal Validation
SPR data confirms electrochemical results, and vice versa. Two independent transduction methods on one interface give higher confidence in every design decision.
From Academia to Diagnostics
Label-free SPR bridges fundamental biosensor research and applied diagnostic development — a single platform that speaks to both communities.

Conclusion
Fontaine et al. establish that label-free SPR provides equivalent analytical performance to electrochemical detection for DNA hybridization — making SPR the rational first step before redox-reporter synthesis begins. Park et al. demonstrate that aptamer-functionalized hydrogel platforms can achieve clinical-grade sensitivity, full reversibility, and weeks of stability in biological fluids. Basini et al. show that MD simulation combined with SPR validation can replace months of SELEX with a targeted computational-experimental pipeline — producing nuclease-resistant PNA aptamers with picomolar affinity for cardiac troponin I. And Garg et al. show what the endpoint of this pipeline looks like: a wearable device that monitors insulin continuously in a living animal, using the same electrochemical aptamer principles that SPR was used to develop and validate.
That translational arc is already reaching commercial scale — Nutromics is bringing the Lab-on-a-Patch® to market, and Aptec Health is moving the Aptalyzer toward clinical insulin monitoring — with benchtop SPR underpinning the optimization work that makes both platforms possible.
The P4SPR sits at the foundation of this pipeline. Probe length, surface density, aptamer conformation, cross-linking geometry, matrix compatibility — all must be optimized before a device can be built. That optimization work is what label-free, real-time, benchtop SPR is built for.
The translation from bench to wearable clinic is happening now, and SPR is how it starts.
References
- N. Fontaine, A. Dauphin, M. Gaida, R. Simard, P. Dauphin-Ducharme, "Electrochemical and plasmonic detection methods yield comparable analytical performance for DNA hybridization," Analyst, 2025, 150, 4389–4394. DOI: 10.1039/D5AN00741K
- S. Park, A. Gerber, C. Santa, G. Aktug, B. Hengerer, H.A. Clark, U. Jonas, J. Dostálek, K. Sergelen, "Molecularly Responsive Aptamer-Functionalized Hydrogel for Continuous Plasmonic Biomonitoring," J. Am. Chem. Soc., 2025, 147, 11485–11500. DOI: 10.1021/jacs.5c01718
- M. Basini, M. Hambli, S. Sahu, R. Corradini, "Rational design of PNA aptamers for cardiac troponin I detection," Nano Today, 2026, 67, 102957. DOI: 10.1016/j.nantod.2025.102957
- M. Garg, S. Chupil, J. L'Heureux-Hache, R.A. Lal, M. Poudineh, L. Soleymani, "Integrated hydrogel microneedle aptamer sensor for wearable insulin monitoring," Nature Reviews Bioengineering, 2025. DOI: 10.1038/s44222-025-00394-3
- E.M. McConnell, M. McKeague, A. Lin, P. Mallikaratchy, R. Patel, S. Shigdar, M.C. DeRosa, "In vitro selection of aptamers and their applications," Nat. Rev. Methods Primers, 2023, 3, 54. DOI: 10.1038/s43586-023-00238-7
- Nutromics. Lab-on-a-Patch® — Continuous Diagnostic Monitoring. Available: nutromics.com
- Affinité Instruments. Available: affiniteinstruments.com
Acknowledgements: Findings in this note are based on published work of Fontaine et al. (2025), Dauphin-Ducharme group, Université de Sherbrooke; Park et al. (2025), BioMed X Institute and collaborators; Basini et al. (2026), University of Parma; and Garg, Poudineh, Soleymani et al. (2025), McMaster University / University of Waterloo / Aptec Health.
TN-02 — Affinité Instruments Technical Note Series